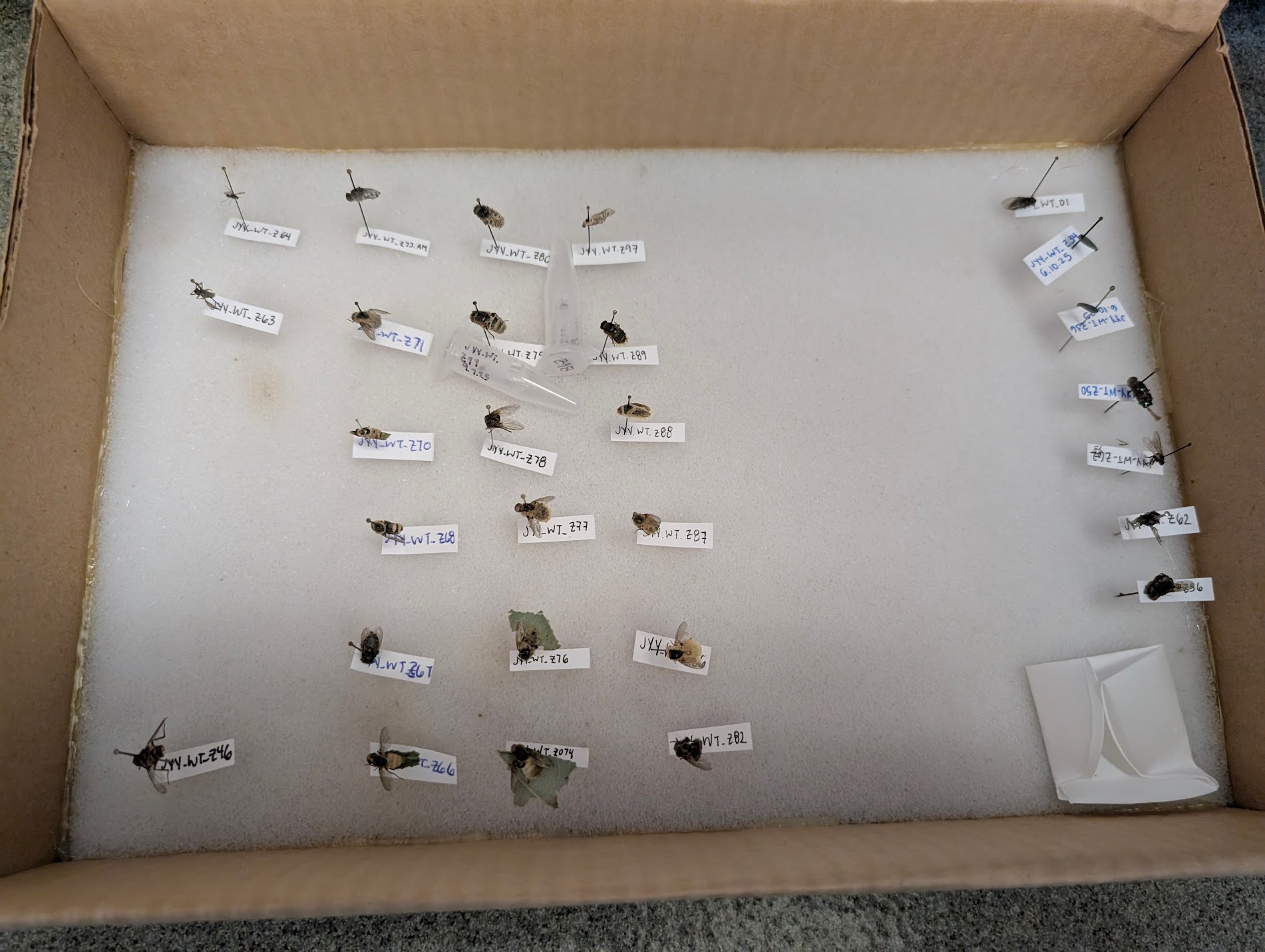
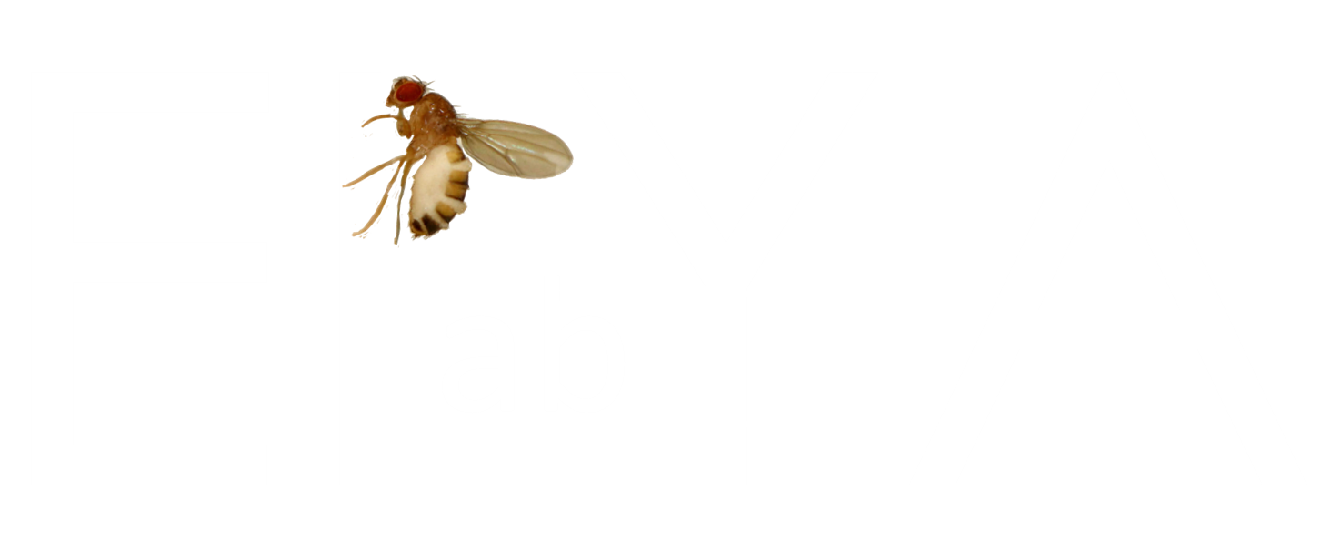
Here’s the process we apply for each sample that enters our laboratory:
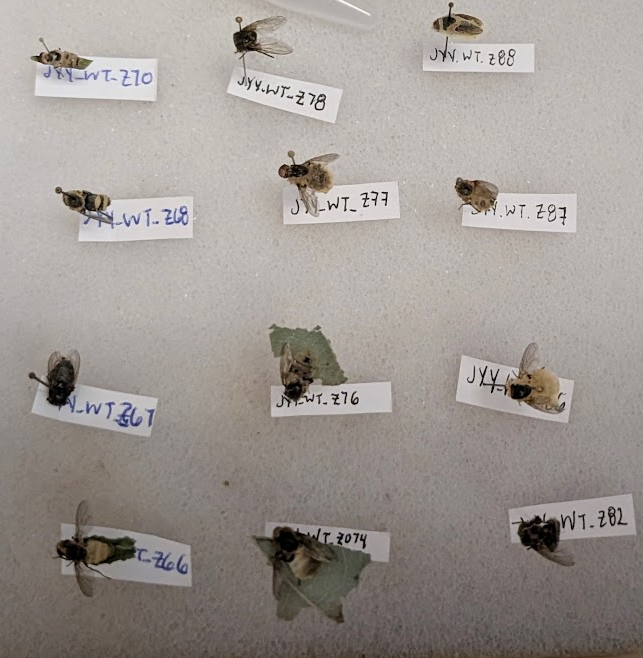

- Receive specimen.
- Inspect specimen under microscope.
- If specimen is sufficiently robust, pin.
- Take high-resolution images at Museum of Comparative Zoology.
- Biopsy fungal and host tissue and extract DNA.
- Quantify DNA recovered.
- Amplify diagnostic loci (host: COI, fungus: LSU)
- Confirm amplification by gel electrophoresis.
- Prepare minION sequencing library.
- Sequence library.
- Use custom computational pipeline to find consensus sequences among reads.
- Search NCBI databases for best matches.
- Assign ID based off these matches.
- Add sequences to growing phylogenetic trees.


This pipeline was developed and is executed solo by postbac Jessenia Yupangui Yupa.

